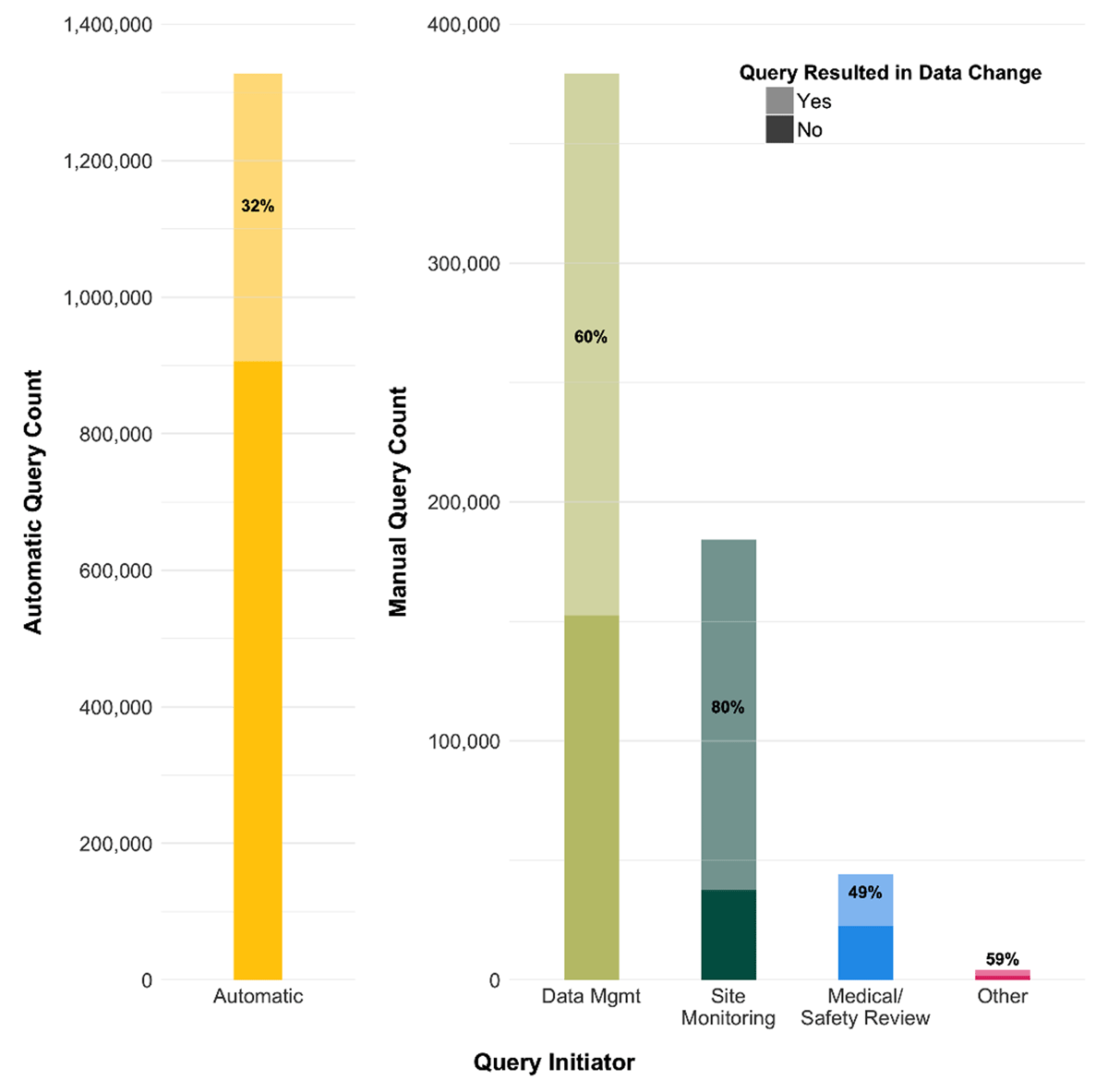Introduction
Over the last years, regulators encouraged sponsor companies to follow a risk-based approach to quality management (RBQM) and the maxim became “focus on what matters most”1. While many sponsor companies and contract research organizations (CROs) were already moving towards a more targeted RBQM approach, the recent COVID-19 pandemic has catalyzed this thinking on a broad scale. The pandemic poses numerous challenges to clinical trial programs. For example, due to lockdown, social distancing, and clinical sites working in crisis mode, visits per study subject decreased nearly 30% between October 2019 and April 2020.2 Sponsors are working to lessen the potential burden on study sites, and one area under scrutiny is whether a nonspecific query process contributes meaningfully to achieving reliable study conclusions.
A pivotal literature overview by Sheetz et al. determined that even though SDV consumed a significant amount of effort, it resulted in just 1.1% of data being updated.3 This observation led many sponsors and CROs to adopt targeted SDV approaches, guided by site performance indicators and data criticality. The latter is defined as data that are used to make decisions about the investigational product’s safety and efficacy profile.4 Although SDV still corrects critical transcription errors and provides information on the accuracy of data entry, the risk-based approach shifts the focus to more strategic quality management methods in site monitoring, such as source data review and centralized monitoring. This example of the reconsideration given to the relevance of one data quality approach helped motivate the current study of the generalized query process.
The current query process consumes time and resources from both the sponsor and the clinical research site. Automatic queries must be specified, programmed, and tested and manual queries must be defined by the Clinical Data Manager, the Study Medical Expert, the Drug Safety Specialist, or the Site Monitor. Manual queries and automatic queries that are triggered once data is entered then require review and response by site personnel and subsequent review and action again at the sponsor.
The cost of queries is not known precisely, with estimates varying from $28 to $225 per query.5,6,7 However, with approximately 2,000 Phase III studies initiated annually – each generating tens of thousands of queries – it is clear that an inordinate amount of resources is spent industry-wide on query generation and resolution. Since there is no guidance stipulating the details of the query management process, this is largely a self-imposed activity.
Earlier publications explored various approaches to risk-based quality management in CDM. In a relatively small study analyzing the effectiveness of 1,397 queries, it was shown that queries led to a change of only 2 percent of the data,8 supporting recommendations to revisit the query management process.9,10
Moreover, since the untargeted query approach is essentially at odds with a risk-based (i.e., targeted) approach to quality management, an inquiry into the utility of the query process is justified.
To confirm the earlier findings and to explore the extent to which the traditional query process contributes meaningfully to achieving reliable study conclusions, clinical data management representatives of seven pharmaceutical companies teamed up with a clinical trials solution vendor to analyze a collective data pool. To assess the value of the generalized query process, the following questions were asked: what functions are initiating the queries, how are the queries distributed over various form types, and how frequently do queries result in a data modification?
Methods
Study and Data Selection
Twenty Phase III studies with primary endpoint completion dates since 2014 were randomly selected from the Medidata Enterprise Data Store to represent 5 therapeutic areas (TAs). Eligible studies originated from the participating companies, supplemented with studies from other sponsors with appropriate data rights for use of de-identified operational data. The query records corresponding to these studies were extracted to a secure environment for analysis.
Data Standardization
Since the data originated from different companies, there was no universal terminology identifying form/domain names or field/variable names. Medidata had previously developed a semi-supervised machine learning Form and Field Classifier (FFC) model,11 which was applied to the combined query data to generate a standardized form and field mapping for each query record. Suggested FFC mappings were independently reviewed by 2 subject matter experts, with final consolidation of discrepant judgements by a 3rd individual.
Each query record was classified as to whether the query resulted in a change or not, and whether a change was direct or indirect. As shown in the examples in Table 1, a change was classified as direct if the data point that was updated was the same as the one that was queried. An indirect change was defined as any other modification propagated by the query process, including a change to a different data point than that queried, or to a source record associated with the study subject. Direct changes were detected by comparing the original data value queried to the answered data value of the closed query. Indirect changes were identified by first evaluating word maps of the free form text response for those queries that resulted in a direct change. The most common words and phrases that indicated a direct modification had occurred (e.g., “updated”, “data corrected”, “added”) were then used to identify those queries that led to an indirect change.
Examples of queries resulting in direct and Indirect data changes.
| Query Text | Queried Data | Answered Data | Answer Text | Type of Change |
|---|---|---|---|---|
| Age is missing – please complete. | 46 | Data entered | Direct | |
| Age data does not match source. Please verify. | 55 | 55 | Verified source was incorrect. Source was updated. | Indirect |
| Heart rate is recorded as 59 bpm (<60 bpm). Please correct the heart rate or report the abnormality “Sinus bradycardia” in the section “Did the subject experience any rhythm abnormality?” | 59 | 59 | Updated as sinus bradycardia | Indirect |
Changes were further categorized as “non-informative” if the query and its response only addressed the format of unknown information. For example, “Unknown Strength” corrected to blank, blank corrected to “Unknown”, or “UN:UN” corrected to “Unknown” were labeled as non-informative data changes.
In the Medidata Rave EDC (electronic data capture) system, each manual query has an associated role assigned to the user initiating the query. In consultation with the CDM network participants, these role names were mapped to one of 4 query initiator roles: Data Management (DM) (e.g., “Site from Data Monitor” and “Site from DM”), Site Monitoring (e.g., “Site from CRA” and “Site from Site Monitor”), Medical/Safety Review (e.g., “Site from Medical Monitor” and “Site from Safety”), or Other (e.g., “Site from Study Team” and “Site from Adjudicator”).
Data Analysis
Counts of data points by standardized form name were generated for all data and counts of standardized field names were created for all queried records. Query Efficacy was defined as the percentage of queries that resulted in a direct or indirect data change. Heat maps of data changes were normalized using the square root of data counts. Data summarization was performed in Python, SAS, and R.
Results
Studies and Data Points
The 20 studies selected for analysis represented 4 studies in each of 5 TAs. The studies encompassed a total of 20,125 study participants and 49,259,945 data points. Study enrollment ranged from 200 to 4200 participants, with a median of 600 participants per study. The study duration to primary endpoint completion ranged from 2–54 months, with a median duration of 29.3 months.
Queries by Therapeutic Area
In total, 1,939,606 queries were generated, 68% (1,327,526) automatically triggered and 32% manually created (612,080), for an overall query rate of 3.9%. The distribution of queries by TA was similar for manual and automatic queries.
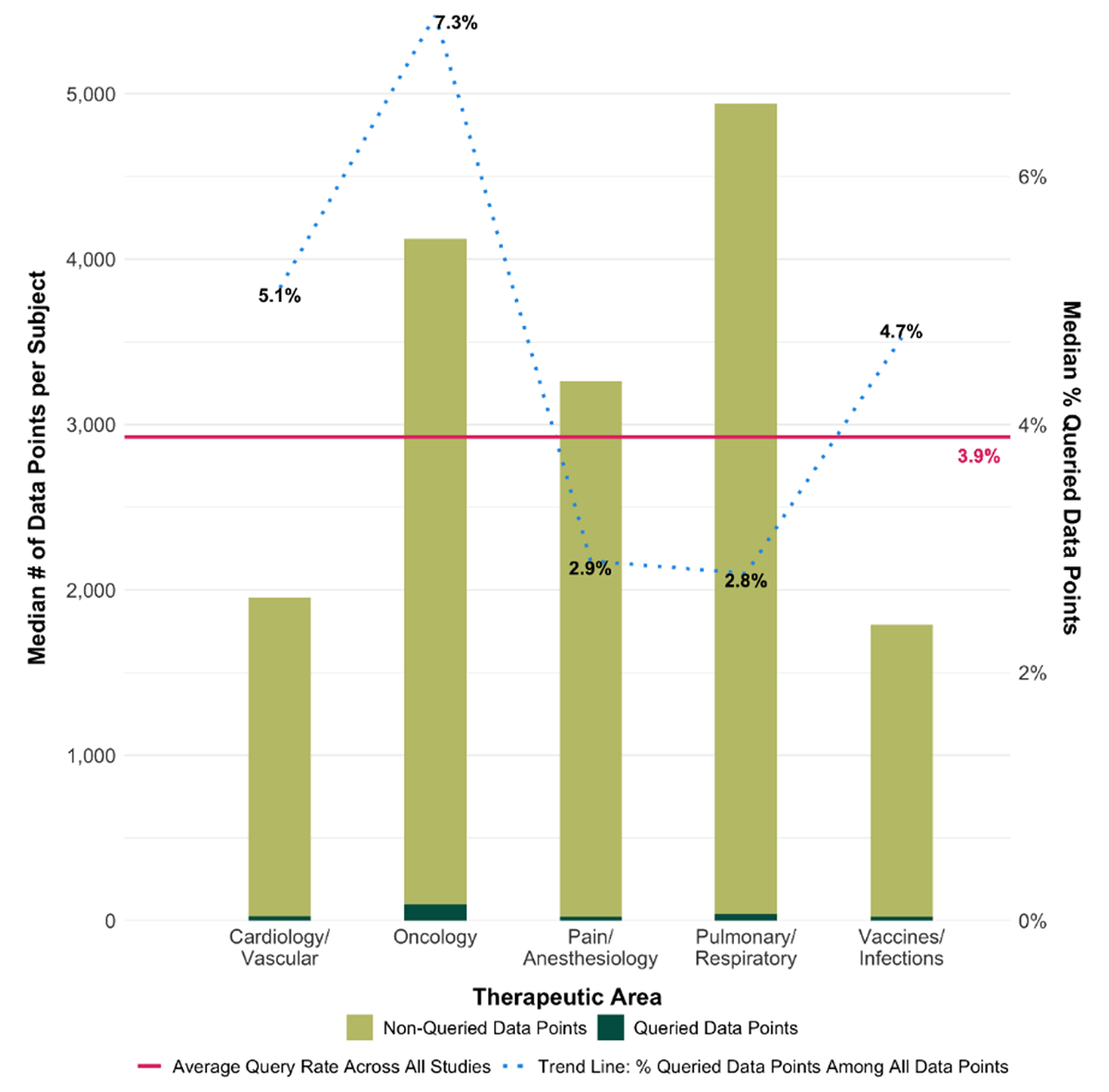
The number of queries per study averaged 96,980, with a median query rate (number of queries per total number of data points), ranging from 2.8% for Pulmonary/Respiratory studies to 7.3% for oncology studies, as shown in Figure 1.
Queried and non-queried data points by therapeutic area.
The left y-axis corresponds to the median number of data points collected per subject and the right y-axis to the median percentage of data queried. The x-axis depicts the relative proportion of queried and non-queried data per TA. The dashed trend line corresponds to the right axis and shows the median percentage of queried data points by TA, and the solid line represents the overall proportion of all data queried.
Queries by Form Type
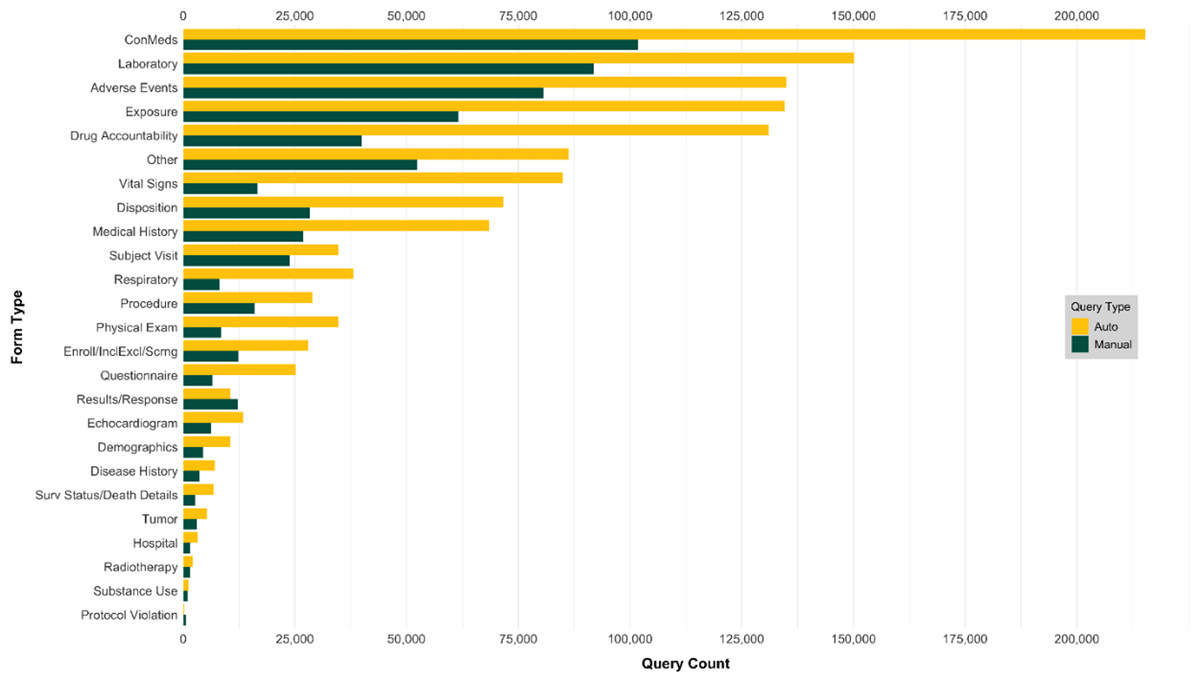
The 1,939,606 queries originated from 1,238 unique form names and 8,461 unique field names. Using the FFC model, the queries were mapped to 25 standardized form types and 196 standardized fields. As shown in Figure 2, five forms, representing those typically collected throughout a study (ConMeds, Laboratory, Adverse Events, Exposure, and Drug Accountability), accounted for more than half of all queries (61% of manual queries and 57% of automatic queries).
Queries by Initiator
There were 29 unique role names assigned to users who entered queries in the 20 studies in this analysis, ranging from 2 to 7 (median = 4) roles per study. DM initiated 62% of the manual queries, with Site Monitoring representing the next largest initiator role (30%). Queries by Medical/Safety Review were more common in Oncology (12%) and Cardiology/Vascular (8%) studies than within the other TAs (Pain/Anesthesiology 5%, Vaccines/Infections 3%, Pulmonary/Respiratory <1%).
Query Efficacy and Data Changes
Queries were generated for 3.9% of entered data (1,939,606/49,259,945). The overall Query Efficacy was 42%, with 818,921 of the 1,939,606 queries resulting in either a direct (n = 697,185, 85.1%) or indirect (121,736, 14.9%) change. Only 4% of indirect changes involved modifications made to source data. Automatic queries were less efficient than manual queries with a Query Efficacy of 32% (421,339/1,327,526) vs 65% (397,582/612,080). Requeries represented 10% of manual queries.
Overall, the sum of automatic and manual queries led to 818,921 changes in the database, which pertain to 1.7% of the data points. Automatic queries led to a change of 421,339 data points out of a total of 49,259,945 (0.9%), whereas manual queries led to a change of 397,582 out of 49,259,945 data points (0.8%).
Three percent (11,086) of manual queries and 2% (8,386) of automatic queries resulted in a data change that was categorized as non-informative.
There is wide variability in Query Efficacy by initiating function as shown in Figure 3. SDV/Site Monitor-initiated queries had a higher average Query Efficacy (80%) than other manually- created queries. Queries initiated by DM were more likely to result in a data modification (60%) than the overall Query Efficacy of 42%, but slightly below the average 65% Query Efficacy for manually generated queries. Queries generated by Medical/Safety Review resulted in the fewest data changes with a Query Efficacy less than 50%.
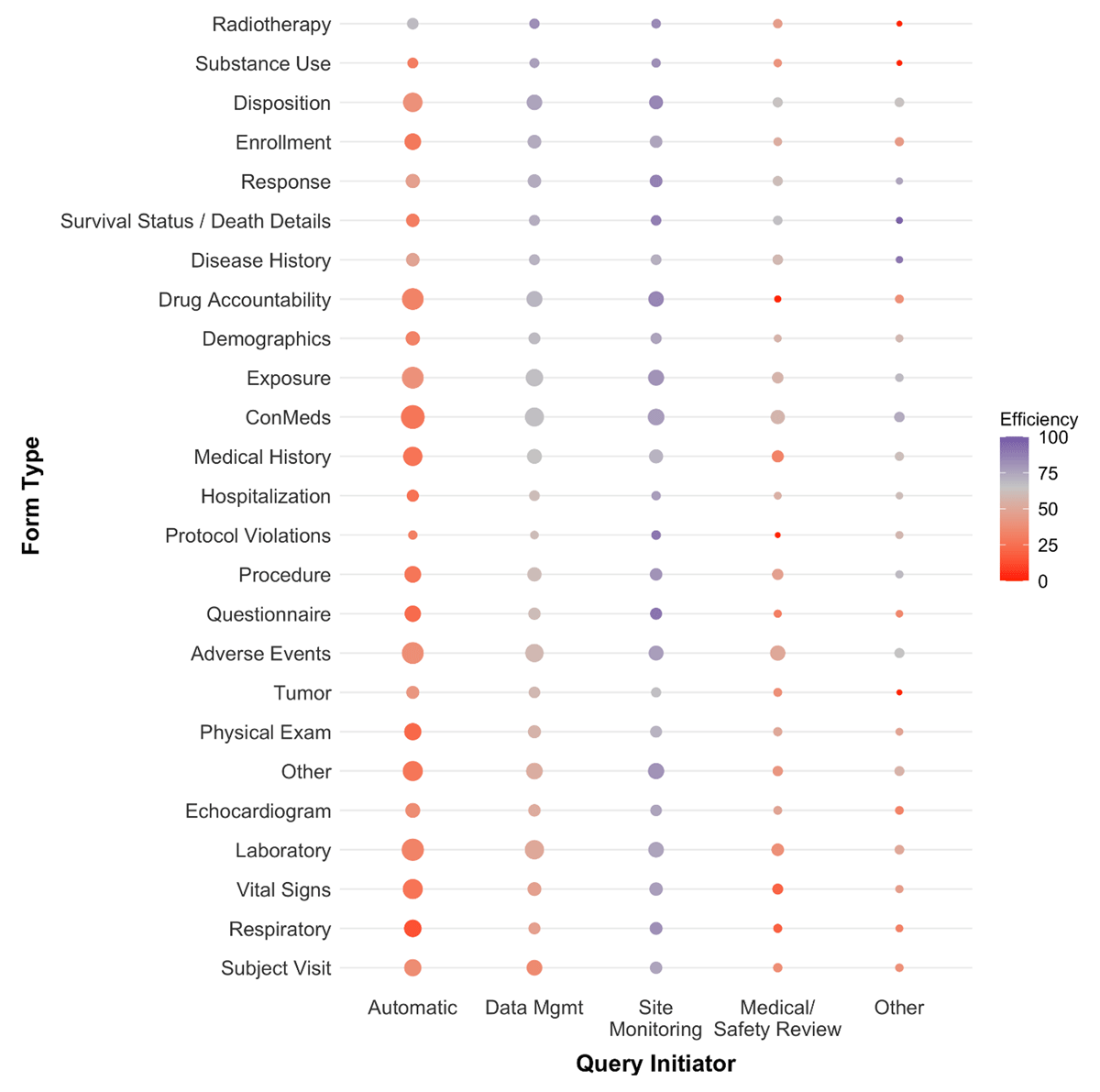
When Query Efficacy is evaluated by form type, the general change pattern by initiating function is observed but some notable patterns emerge. The heat map shown in Figure 4 indicates where the different functions provide the most efficacious contribution, and where their contribution is of limited value. Considering the low overall effectiveness of the query process, focusing on critical form types only seems to be the most productive strategy.
Query Efficacy heat map per form type and initiating function.
The bubble size represents the relative proportion of queries, and the strength of the bubble color represents the degree of Query Efficacy, ranging from red (low efficacy) to purple (high efficacy). The neutral gray Query Efficacy color is centered at the average manual Query Efficacy rate of 60%. Form types are sorted sequentially from most to least effective queries based on those initiated by DM. For example, within each Query Initiator function Concomitant Medications (ConMeds) represent form types that are queried more frequently (larger bubbles). Automatically generated queries for ConMed form types were among the least efficient queries (darker red color), but had higher than average Query Efficacy when initiated by Site Monitoring (darker purple color).
Our analysis found that only a small proportion of the 49,259,945 data points is ever queried, and that on average these queries lead to a data change less than half of the time (42%).
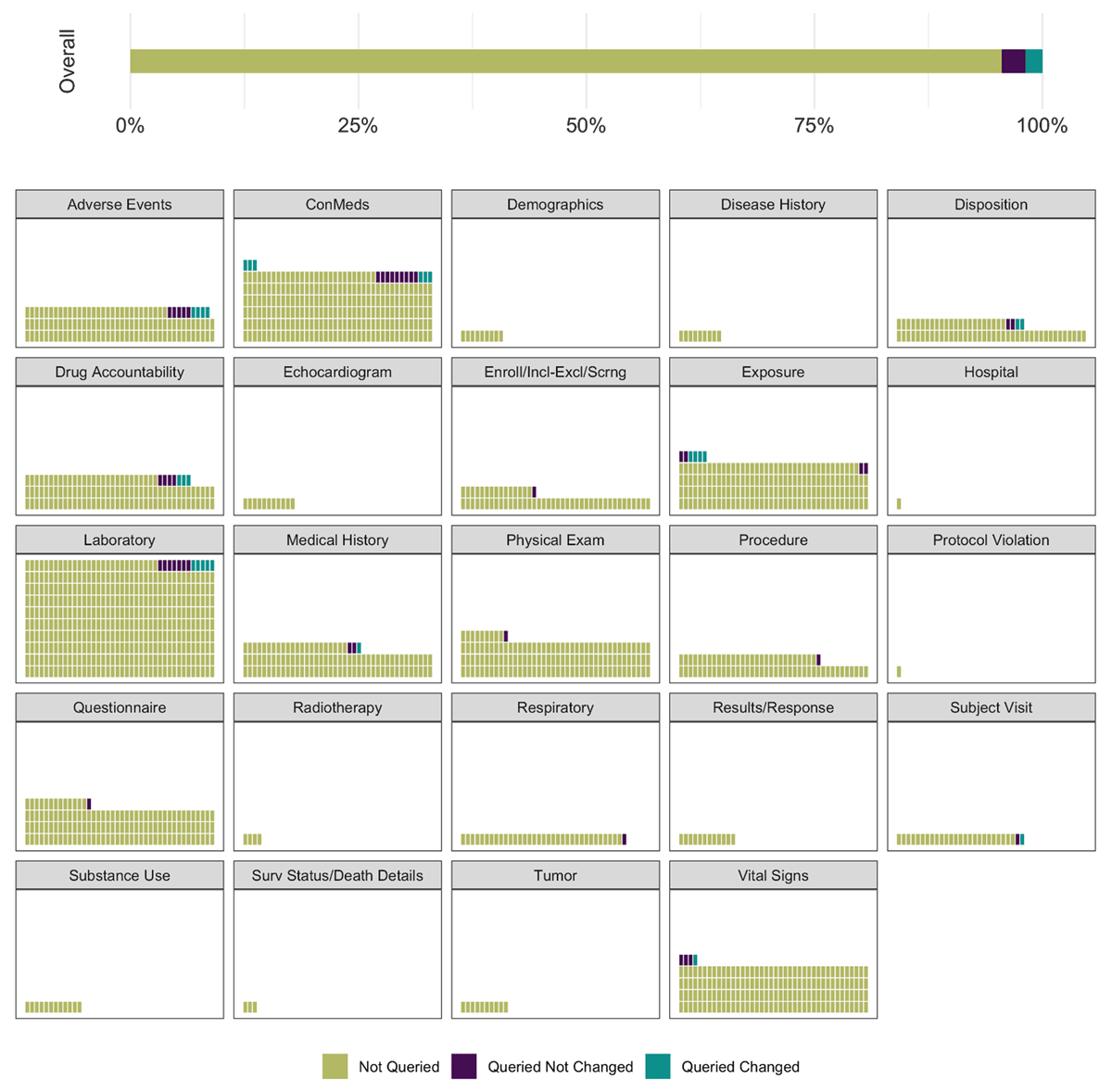
Thus, as visualized in Figure 5, the majority (>98.3%) of data collected in a study is never changed as a result of the query process.
Distribution of data not queried, queried but not changed, and queried and changed, overall and by form type.
The individual waffle plots represent the total number of data points of each specific form type, standardized to Laboratory forms. Thus, each block of the waffle plots represents 0.25% scaled to the Laboratory form type.
Discussion
Collecting data in clinical studies started as a very manual process: transcribing data onto the paper case report forms (CRF) and double-data entering this information into the clinical data management database. In this error-prone process, a culture of check, double check, and retrospective data correction evolved. However, since these early paper days, the intrinsic quality of the data acquisition process has dramatically increased12 – with an associated diminishing efficacy of the retrospective cleaning process. The current trend towards a prospective focus on quality by design, and centralized data review will further reduce the value of retrospective data cleaning.
The combined quality of the current process steps leading up to the data in the database – including protocol, CRF and database design, a comprehensive body of front-end edit checks, monitoring strategy, and site training – leads to a highly accurate set of data, witnessed by the fact that we found that only 3.9% of all data was ever queried and that less than 1.7% of all entered data was changed during the query process. Previous work8 focused on changes made to the EDC database, which can directly impact study results. Our analysis encompassed both these direct, as well as indirect changes that result from the query process in order to fully appreciate impact of the query process on overall data quality. Our study confirms the findings of previous authors that demonstrated that the current process is not efficient.
Furthermore, in our study cohort, the overall Query Efficacy of 42% was shown to depend on the initiating function and the form type. Forms that are less likely to result in a data change and that have a low criticality likely indicate areas where time and resources could be more effectively targeted. From this we have concluded that functions could put the focus on the most critical data and on domains where it is most efficient.
Our results are similar to those reported previously by Sheetz et al. Whilst evaluating the value of SDV as quality control measure in Clinical Trials, this retrospective data analysis based on a massive amount of 1168 Phase 1–4 studies found that the percentage of data corrected by automatic queries was 1.4%, whereas the portion of non-SDV data corrections initiated by data management, medical, safety, and biostatistics amounted to 1.2%.3
In our sample of 20 Phase III studies, nearly two million queries were generated, on average 96,980 queries per study. Assuming the costs per query to be somewhere between $28 and $225, we can estimate that companies spent anywhere from 2.7 to 22 million dollars per study to clean less than 1.7% of the data. Errors can be cleaned more efficiently and systematically by means of centralized tools that detect outliers, unusual data trends, systematic errors, and other data integrity problems. The very resource-intensive query-based data cleaning can be limited to the most critical data (‘targeted data cleaning’). Remaining errors are likely to be infrequent, and assuming that most mistakes occur at random and are unlikely to substantially impact analyses, any benefit in cleaning them in a large enough study is doubtful.12
We also observed situations (2.4% of all queries) in which the reason for the query was simply to ensure that unknown information was entered in the right format. Clarifying expectations for recording values not collected, ideally at an industry-wide level, would prevent these immaterial queries.
Improvements to the current query and data collection process may be achieved by
Focusing on critical data by eliminating questions and assessments that do not contribute to answering the scientific question of the protocol
Clearly articulating queries and specifying expectations for recording values that are unknown
Limiting queries that ask the site to confirm information
Further improving the quality of automated checks to increase its Query Efficacy of 32%
Improving the quality of the site training and data entry instructions
Putting more emphasis on centralized data review to assess completeness and to identify outliers, anomalies, and patterns at study, site, and country level
There are several limitations of this work. While the machine learning predictions of the mapped form type and field name were reviewed by subject matter experts, errors in the final classifications may exist. Additionally, a large number of forms were classified as “Other” indicating an opportunity to further refine the model with additional standardized form and field targets. The classification of indirect data changes was informed by the verbiage of queries resulting in a direct change. While this was the most expedient method of identifying such changes described in free-form text, we recognize that it may not be fully accurate. Finally, the focus on Phase III studies may not reflect the query process in other study phases, nor may the therapeutic areas selected for this project be representative of all TAs.
Despite these shortcomings, we feel that this study offers important insights towards improving the query management process. Not only have we confirmed that only a very small proportion of data is ever queried and changed, this work identifies concrete steps towards achieving a risk-based approach to query management. To our knowledge, this is the first large-scale study to quantify the efficacy of the manual and automatic query process and to assess direct, indirect, and non-informative data changes. This work was largely made possible by the ability to apply a machine learning approach to map form and field names to a standard vocabulary. While this work commenced well in advance of the COVID-19 pandemic, its findings seem particularly timely as companies currently are forced to triage the most relevant data in the pandemic-affected data flow.
Conclusions & Recommendations
Our findings are consistent with, and expand upon, previous studies that determined that data cleaning results in a minimal number of changes to the underlying database. Even including indirect and non-informative changes, we determined that only 3.9% of data is ever queried, and that only 42% of these queries resulted in a data modification, affecting less than 1.7% of all entered data. We also observed that Query Efficacy depends on the type of query, type of form and query initiator.
Our recommendations include the following:
Ending the generalized use of automated and manual queries of non-critical form types and fields to correct database errors in Phase 3 studies. This builds on previous suggestions8 and is a rational step to end a resource-intense activity with a very low added value since, by definition, these data are not directly relevant to the safety and well-being of the subject, nor the scientific validity of the study.4
Increasing the use of Query Efficacy metrics for critical form types and fields – in combination with pre-specified Risk and Quality Tolerance Limits – to identify systemic issues in the data collection process.8
Combining Query Efficacy metrics with additional quality indicators – such as query rates per site and CRF, SDV metrics, and the number of requeries – to support quality by design (QBD) approaches. QBD targets may include protocols, CRFs, training materials, user guides, and query wording.
Unambiguously specifying expectations for recording values that are unknown, to prevent non-informative queries and irrelevant data updates.
Incorporating the use of augmented intelligence tools to supplement human review of data.
For these recommendations to be viable, Clinical Data Management, other internal sponsor reviewers of the data (Medical Review, Statistics, Regulatory), GCP inspectors, and health agency reviewers will have to embrace that quality data is not necessarily error-free data, but rather fit-for-purpose data that sufficiently supports conclusions equivalent to those derived from error-free data.13 With the typical clinical study collecting hundreds to thousands of data values for each subject, a minimal occurrence of random errors and omissions in non-critical data would be inconsequential and not indicative of systemic quality issues.
Only with high-level internal and external support for this risk-based approach to quality management within CDM – endorsed and promoted by regulators and ICH E6(R2)14 and ICH E8(R1)15 – will we be able to change the wasteful culture in which sponsors and sites feel it as their duty to spend a disproportionate amount of time creating and responding to queries affecting a quantitatively and qualitatively minimal portion of the data, instead of focusing on what matters most.
Competing Interests
The authors have no competing interests to declare.
References
1. European Medicines Agency. Reflection paper on risk based quality management in clinical trials. 2013. EMA/269011/2013.
2. Medidata Solutions, Inc. COVID-19 and Clinical Trials: The Medidata Perspective, Release 5. 2020. Available at https://www.medidata.com/wp-content/uploads/2020/05/COVID19-Response5.0_Clinical-Trials_20200518_v2.2.pdf. Accessed June 12, 2020.
3. Sheetz N, Wilson B, Benedict J, et al. Evaluating Source Data Verification as a Quality Control Measure in Clinical Trials. Ther Innov Regul Sci. 2014: 48(6): 671–680. DOI: http://doi.org/10.1177/2168479014554400
4. TransCelerate Biopharma Inc. Position Paper: Risk-Based Monitoring Methodology. 2013. Available at https://www.transceleratebiopharmainc.com/wp-content/uploads/2013/10/TransCelerate-RBM-Position-Paper-FINAL-30MAY2013.pdf
5. Lane J. Data Collection Mistakes: A Cost Driver for Poorly Performing Clinical Sites. Longboat; 2018. Available at https://blog.longboat.com/data-collection-mistakes-a-cost-driver-for-poorly-performing-clinical-sites. Accessed April 23, 2020.
6. Law S. Clinical Data Quality & Query Resolution Costs. Linkedin; 2017. Available at https://www.linkedin.com/pulse/clinical-data-quality-query-resolution-costs-steven-law/. Accessed April 23, 2020.
7. Ensign L. Improving the Quality of Queries in an RBQM World. In: 23rd DIA Japan Annual Workshop for Clinical Data Management. 2020; 124–130. Tokyo: DIA.
8. Mitchel J, Tantsyura V, Kim Y, Timothy C, McCanless Dunn I, Nusser-Meany M. Query effectiveness in light of risk-based monitoring. Data Basics. 2018; 24(4): 4–10.
9. Tantsyura V, Mitchel J, Kim Y, et al. Impact on data management of the new definitions of data quality (DQ), risk-based approaches to quality and eSource methodologies. Data Basics. 2016; 22(5): 4–23.
10. Tantsyura V, Mitchel J, Kim Y, Mijaji T, Yamaguchi T, McCanless Dunn I. Risk-based approaches to data management and data quality: double standards or is it just common sense and intelligence taking over? Data Basics. 2017; 23(1): 13–16.
11. Zhang FEM. Automate an Extended-SDTM Mapping in Clinical Trials using Machine Learning. In: EU Connect. 2018; e1–16. Frankfurt: Pharmaceutical Users Software Exchange. Available at https://www.phuse.eu/euconnect18-presentations.
12. Mitchel JT, Kim YJ, Choi J, et al. Evaluation of data entry errors and data changes to an electronic data capture clinical trial database. Drug Inf J. 2011; 45(4): 421–430. DOI: http://doi.org/10.1177/009286151104500404
13. Meeker-O’Connell ABL. Current trends in FDA inspections assessing clinical trial quality: an analysis of CDER’s experience. The Food and Drug Law Institute Update. March/April 2011.
14. Food and Drug Administration, US Department of Health and Human Services. ICH E6(R2) Good Clinical Practice: Integrated Addendum to ICH E6(R1), March 2018. Available at https://www.fda.gov/regulatory-information/search-fda-guidance-documents/e6r2-good-clinical-practice-integrated-addendum-ich-e6r1.
15. Food and Drug Administration, US Department of Health and Human Services. ICH E8(R1) Harmonized Guideline General Considerations for Clinical Studies, Draft version August 2019. Available at https://www.fda.gov/regulatory-information/search-fda-guidance-documents/e8r1-general-considerations-clinical-studies.